View on GitHub
Open this notebook in GitHub to run it yourself
Chebyshev approximation of the inverse function
In this notebook we present the four ways of approximating the inverse function with Chebyshev polynomials:- Using Classiq’s QSP application.
- Using Numpy interpolation with a give degree, here the polynomial values are bounded in .
- Using Numpy interpolation with the theoretical degree, and then trimming to a smaller degree.
- Using an optimized (for the Maximum norm) expansion according to https://arxiv.org/pdf/2507.15537 (available in Classiq QSP application).
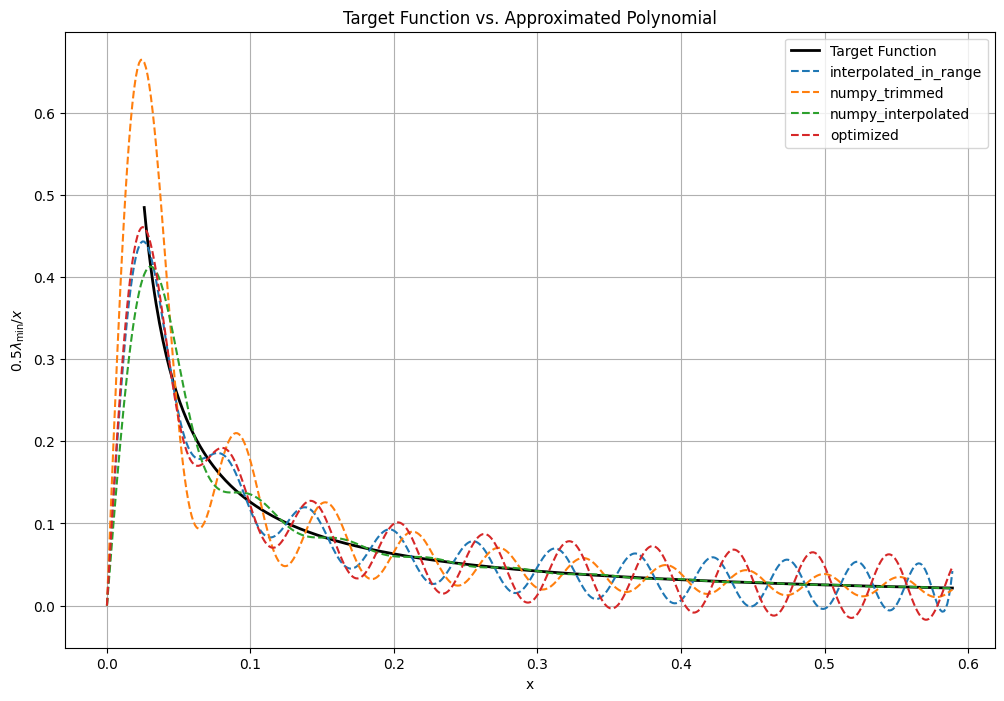
Output: